Continuing from my post on the principles of MeDIP, I'd like to go over the steps in a MeDIP experiment and discuss some optimization tips.
Our lab employs MeDIP-seq for use in cell-free DNA. While we do not do any immediate applications with genomic DNA isolated from tissue, I will describe the general method for genomic DNA and points that should be considered for MeDIP-seq—the most prominent use of MeDIP.
DNA Shearing
While most library generation techniques require fragmentation of the DNA, shearing DNA at a particular size for MeDIP is especially important. The fragmentation of these fragments should range from 200-600 basepairs. This is important because the anti-5mC antibody requires the presence of more than one methylated cytosine for binding—if the fragments are too short then the chance of having multiple methylated cytosines is reduced. However if fragments are too large, then it can be difficult to discern what proportion of that region was methylated.
Once DNA has been sonicated and size-selected, sequencing adapters (typically Illumina or NEB) need to be ligated to the DNA for subsequent steps and analysis. Adapter ligation is typically performed after an End Repair and A-Tailing reaction, to promote ligation.
Adding the Anti-5mC Antibody
A relatively simplistic step. However there are several points that should be considered when adding the anti-5mC antibody.
The DNA that has now been ligated with sequencing adapters is denatured at 95 degrees (C) for 10 minutes. Our lab uses a standard PCR thermocycler to achieve this. It is important to immediately remove the tubes and transfers to an ice water bath after 10 minutes have passed. However, depending on the PCR tubes, the caps may open due to built up pressure—therefore it is important to ensure that the caps are closed immediately to prevent evaporation of sample. While typical protocols do not emphasize on the duration of adding the antibody after placing the samples in ice-water, we have noticed if the antibody is added 20-30 minutes after placing the samples on ice that antibody binding efficiency is severely reduced. Therefore, I recommend that the antibody should be added immediately after the samples have had time to cool (~10 minutes), as this will prevent the DNA from reannealing.
Isolating the Methylated DNA
Magnetic beads are added to the reaction and incubated at 4o degrees (C) for 4-18 hours. These magnetic beads bind to the Fc region of the antibody which is bound to the methylated DNA fragments. After incubation, the samples are placed on a magnetic rack, and the supernatant (which contains unbound and unmethylated DNA) is discarded. Bound DNA is washed to ensure complete removal of unbound DNA and then eluted.

The magnetic rack used in my lab
Assessing Methylation Recovery
While the antibody should ideally recognize methylated cytosine residues only, some non-specific binding does occur. One way of determining this is by adding spiked-in methylated and unmethylated DNA of unique sequences into the MeDIP reaction prior to immunoprecipitation. qPCR can be performed for these fragments, comparing the relative number of methylated and unmethylated DNA fragments in the MeDIP input and MeDIP after immunoprecipitation.
Ideally, the input should show similar amplification for both the methylated and unmethylated fragments, whereas the MeDIP reaction after immunoprecipitation should show a significantly earlier amplification of methylated fragments. Using this approach, you can infer the methylation recovery (IP ct / Input ct) and specificity (typically 99%).
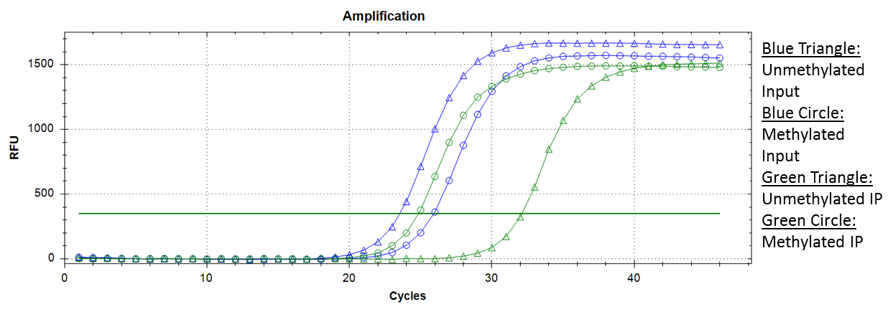
Sample amplification data from a MeDIP experiment
PCR Amplification of the MeDIP-seq Library
Once the DNA library has been enriched for methylated DNA, the final step is performing a PCR amplification reaction with primers targeting regions of the library sequencing adapters previously ligated. This results in an amplification of fragments that can be sequenced. Depending on the amount of input DNA used, it may be necessary to do a qPCR reaction to 30-40 cycles (with 1 uL DNA) to determine the optimal number of PCR cycles. PCR overamplification should be minimized to ensure superior quality of the library.
The optimal cycle is typically where DNA concentrations begins to exponentially increase, before the linear phase and plateau. Once the optimal cycle has been determined and the PCR reaction has completely, DNA can be isolated and size-selected by any preferred method (i.e. AMPure beads, gel purification) and submitted for sequencing.
I hope you have found this short series on MeDIP useful, and let me know in the comment section below if you have any questions!
If you're planning a MeDIP experiment, conduct a search on BenchSci to review published MeDIP data using our technique filter to find out which antibody would be suitable for your experimental context.