Ever been frustrated failing to reproduce a key study? Well, unfortunately you’re not alone. In fact, a recent Nature survey found that approximately 50% of biomedical researchers were unable to reproduce their own, or someone else’s, findings. Research antibodies, in particular, contribute much to this irreproducibility.
To address this "reproducibility crisis" in biomedical research, we hosted a symposium at the University of Toronto on Aug 16, 2017, bringing together key opinion leaders and the biomedical research community to discuss how to ensure more reproducible research. A common subject speakers addressed was the importance of antibody validation.
In this article, I’m going to summarize three distinct antibody validation approaches they proposed to help improve reproducibility in biomedical research. We have also made videos from the symposium available on YouTube, and you can find them here:
Approach 1: Standardization
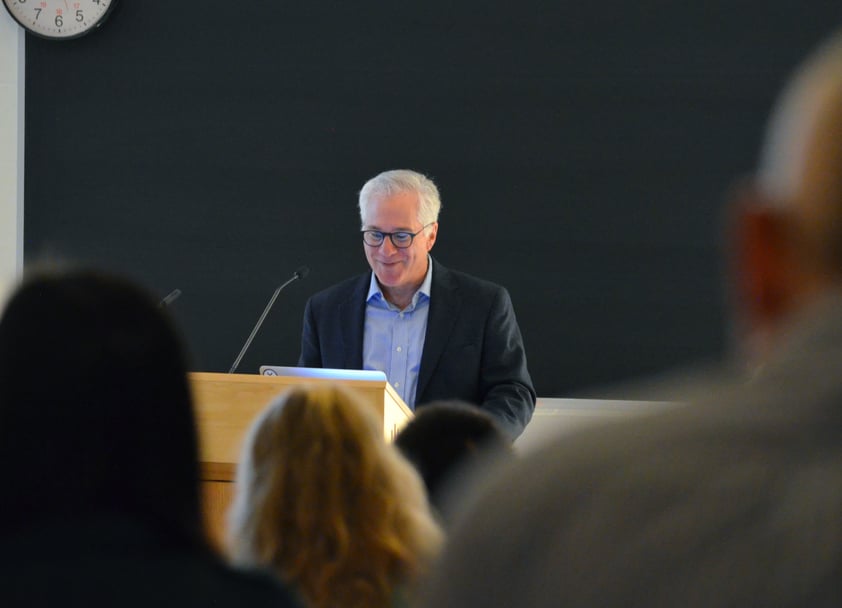

"Standards, when applied the right way, can facilitate the alignment of best practices, reduce variance, and ultimately improve reproducibility"—Dr. Leonard Freedman, President of Global Biological Standards Institute (GBSI)
While industry labs have well-established quality systems for antibody validation, such quality control and assurance frameworks don’t yet exist in academic labs. Therefore, Dr. Leonard Freedman and his organization, the Global Biological Standards Institute (GBSI), advocate for a regulatory approach and lead the development of uniform standards for antibodies within academic research.
Together with funders, publishers, academic scientists and antibody suppliers, GBSI established the Antibody Validation Working Groups to create guidelines and a point system that academic research labs can use to demonstrate and confirm validation of antibodies. Specifically, the guidelines aim to:
- Establish the standards to validate antibodies in applications-specific manner;
- Ensure reproducibility of antibodies over time between manufacturers and lots; and
- Provide training for best lab practices
While it will require some effort, establishing and adopting such standards would ensure that antibodies are uniformly validated prior to publication, thereby improving reproducibility of the findings.
Watch the full presentation by Dr. Freedman here:
Approach 2: Reproducible Reagents
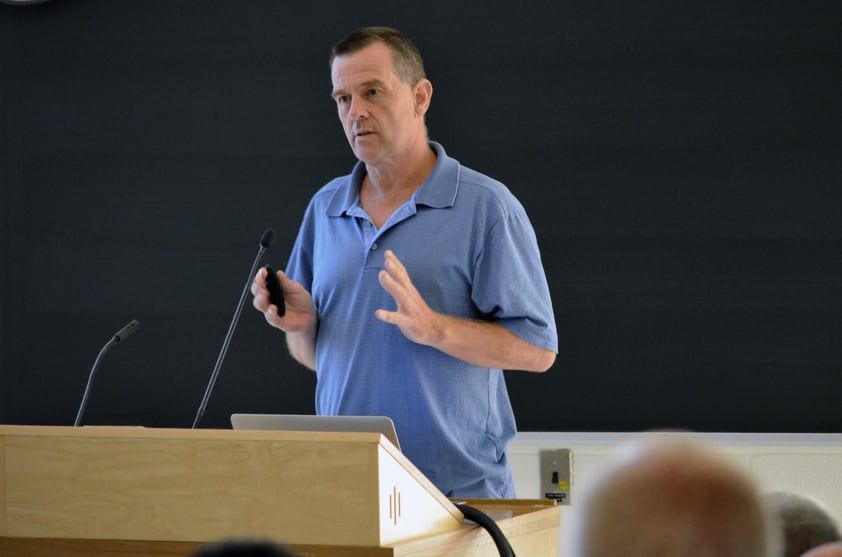

"Reproducibility in biomedical research is impossible until we develop and implement quality metrics for research reagents."—Dr. Aled Edwards, President of Structural Genomic Consortium (SGC)
Dr. Aled Edwards of the Structural Genomic Consortium (SGC) proposed an experimental approach to validate antibodies and improve reproducibility. He argued that the key points to reproducible results are:
- Antibodies themselves must also be reproducible;
- Antibodies must be validated using a quantifiable method; and
- An independent assessor is needed to verify validations
Reproducible antibodies are those that were produced in vitro via phage vectors containing the gene sequence of the antibody. These so-called “recombinant antibodies” eliminate the biological variability associated with the in vivo production of mono- and polyclonal antibodies. Dr. Edwards’ group at SGC has produced thousands of recombinant antibodies and validated each antibody using immunoprecipitation followed by mass spectrometry-based analysis to verify the antigen of interest (Marcon et al, 2015 Nature). The validation data were further replicated by 5 independent labs, ensuring the validity of the antibodies.
Dr. Edward’s proposal has a very strong scientific merit. However, due to the shear amount of protein targets available, producing and validating sufficient antibodies to have any beneficial impact on reproducibility would be a prolonged endeavor.
Watch the full presentation by Dr. Edwards here:
Approach 3: Machine Learning
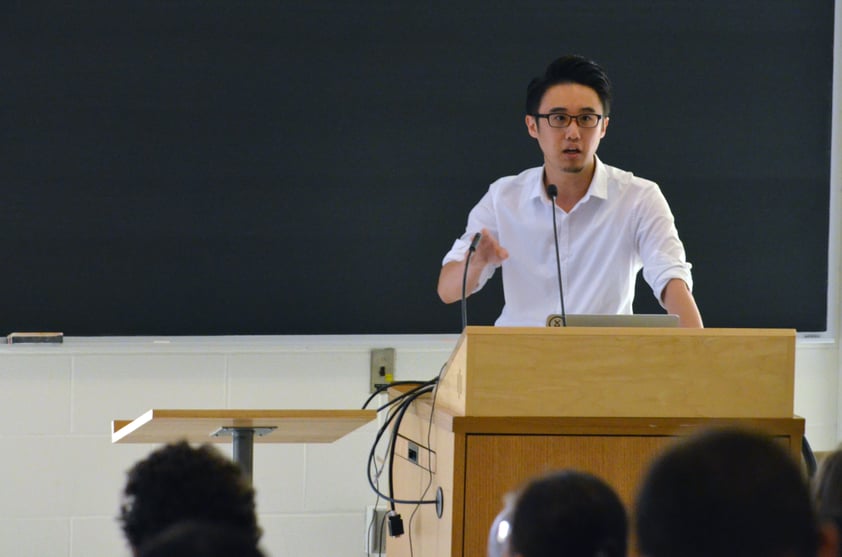

"The most fundamental way to improve research reproducibility is to ensure the reagents are fully and properly cited in your paper."—Dr. Thomas Leung, Chief Scientist at BenchSci
Dr. Thomas Leung, representing our team at BenchSci, talked about our computational and bioinformatical approach to help scientists find validated antibodies and improve reproducibility. The rationale here is that, behind each publication, there is a team of scientists that had validated the antibodies they used, and the identification of such data will help scientists replicate the successful usage of the cited antibodies.
Using a machine learning algorithm, BenchSci analyzes and indexes millions of publications to identify the exact experimental contexts surrounding the use of an antibody. The data produced by the antibody (i.e. the figures) are then curated for users in a manner that is filterable by specific contexts, such as applications, target tissues and cell lines.
Furthermore, BenchSci also captures unpublished results by providing scientists the capability to submit detailed independent antibody reviews. Dr. Leung emphasized how our goal at BenchSci is to provide scientists the data most relevant to their studies, be it published (and analyzed through machine learning) or unpublished (through user reviews), to help them make an informed antibody selection.
Watch the full presentation by Dr. Leung here:
The Take Home Message
The reproducibility crisis is a complex issue that requires a multidimensional solution. Each speaker at the symposium shared with us their own unique perspective on how to improve research reproducibility, with an emphasis on proper antibody validation. Nevertheless, the first step to solving any problem is to ensure that there is sufficient awareness of the problem itself, which is the reason why we hosted the symposium in the first place!
To watch videos from the symposium, head over to our YouTube channel. We would also love to hear your thoughts in the comment section below.